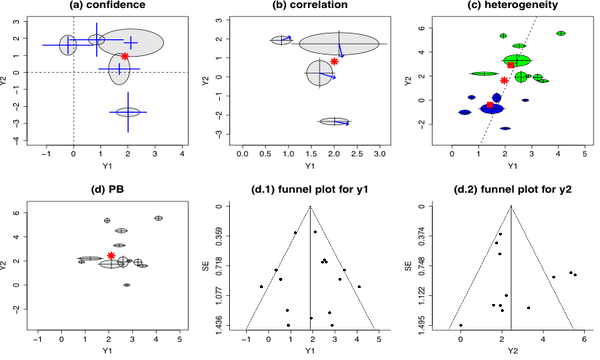
(a). Galaxy-confidence plot:
The galaxy-confidence plot uses cross-hairs to represent the confidence intervals with the cross point showing the point estimate of the bivariate outcome. The confidence intervals are then compared with the lines representing no effect (i.e., black dash lines) to help determine the significance of the effect sizes.
(b). Galaxy-correlation plot:
The galaxy-correlation plot displays the within-study correlation of each individual study by an arrow starting from the center of the ellipse. We restrict the range of arrow to the right side of the ellipse, and the range of correlation (-1, 1) is mapped to the radian range (an arrow above the x-axis represents a positive correlation, while one below the x-axis represents a negative correlation).
(c). Galaxy-heterogeneity plot:
TThe galaxy-heterogeneity plot enables investigations of heterogeneity in a bivariate space. For example, the ellipses in green represent the studies comparing Treatment A and the placebo, while the ellipses in blue represent the studies comparing Treatment B and the placebo.
(d). PB investigation:
In the presence of PB, the galaxy plot is usually not symmetric around the estimated center of mass. More sophisticated statistical methods to quantify and correct for PB can be developed based on this Galaxy plot.